AB024. MIST: a weakly supervised deep learning system for multicenter diagnosis of mediastinal tumors
Graphical Abstract
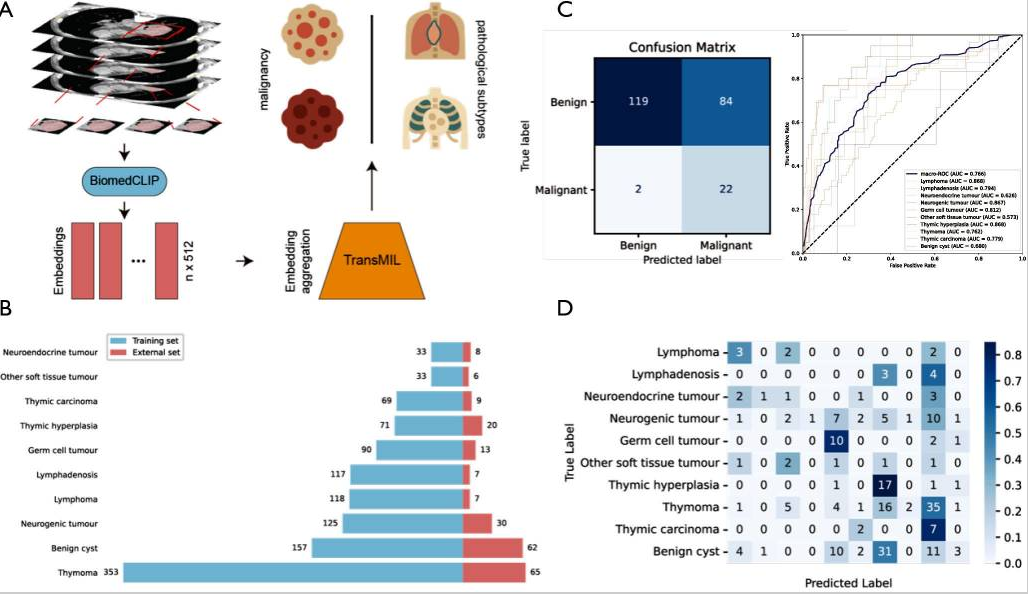
Overview and performance of the MIST system. (A) The overall workflow of the MIST diagnostic system. Slices from chest CT scans are processed by the BiomedCLIP model to extract image features (embeddings). These features are then aggregated by the TransMIL model to predict both malignancy and specific pathological subtypes. (B) The distribution of 10 pathological subtypes across the training set (n=1,173) and the external validation set (n=227). (C) Performance evaluation on the external validation set. The confusion matrix (left) illustrates the performance of binary malignancy classification. The ROC curves (right) show the performance for multiclass subtype classification, with corresponding AUC values for each subtype. (D) A detailed confusion matrix for the 10-class pathological subtype classification on the external validation set. AUC, area under the curve; CT, computed tomography; MIST, Mediastinal tumor Identification with weakly Supervised Training; TransMIL, Transformer based Correlated Multiple Instance Learning; ROC, receiver operating characteristic.
Abstract
Background: Mediastinal tumors represent a distinct group of thoracic diseases, with a rising global incidence and generally poor prognosis. Clinical diagnosis remains challenging due to the complex anatomical structure of the mediastinum and the often ambiguous boundaries between different pathological subtypes. The objective of this study is to develop and validate a high-performance deep learning diagnostic system—Mediastinal tumor Identification with weakly Supervised Training (MIST)—that can accurately and automatically identify mediastinal tumors from chest computed tomography (CT) scans using multicenter data.
Methods: The MIST system was trained using a dataset comprising 1,173 cases of mediastinal tumors with ten pathological subtypes, collected from a high-volume medical center in China. The system processes chest CT scans as input. It first applies a pretrained segmentation model to roughly localize tumor regions within the mediastinum. Then, imaging features are extracted using BiomedCLIP, a multimodal biomedical foundation model. These features are subsequently aggregated by the Transformer based Correlated Multiple Instance Learning (TransMIL) approach to assess tumor malignancy and predict specific pathological subtypes (Figure 1A). MIST’s generalizability was evaluated using an external validation cohort of 227 cases from five independent medical centers (Figure 1B).
Results: MIST demonstrated strong performance in 5-fold cross-validation on the training set, achieving an average area under the curve (AUC) of 0.741±0.045 for malignancy classification and a macro-average AUC of 0.760±0.004 for multiclass subtype classification. In the external validation cohort, MIST maintained comparable performance, with an AUC of 0.818 [95% confidence interval (CI): 0.744, 0.883] for malignancy detection and a macro-average AUC of 0.763 (95% CI: 0.715, 0.807) for subtype classification (Figure 1C). The detailed classification performance is shown in the confusion matrix (Figure 1D). The system achieved a top-1 accuracy of 0.194 (95% CI: 0.145, 0.247) and a top-3 accuracy of 0.480 (95% CI: 0.419, 0.542).
Conclusions: MIST enables accurate identification of both the malignancy and pathological subtypes of mediastinal tumors, with robust generalization across external multicenter datasets. This weakly supervised, foundation model-based approach holds promise for improving diagnostic accuracy and efficiency in clinical settings.
